
-
Pingwei Li
- Professor, AgriLife Faculty Fellow
- Focus Area: structural biology, innate immunity, X-ray crystallography, cryogenic electron microscopy (Cryo-EM)
- Office:
- ILSB 2157
- Email:
- pingwei.li@ag.tamu.edu
- Phone:
- 979-845-1469
Education
- Undergraduate Education
- Northwest University, 1989
- Graduate Education
- Ph.D. Peking University, 1996
- Postdoc. Fred Hutchinson Cancer Research Center, 2001; California Institute of Technology, 2005
Areas of Expertise
- structural biology
- innate immunity
- X-ray crystallography
- cryogenic electron microscopy (Cryo-EM)
Professional Summary
Structural biology and innate immunity
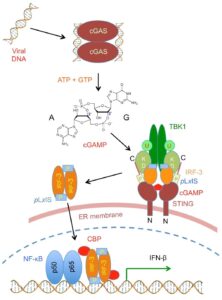
The cGAS-STING pathway in innate immunity.
See Nature 569, 718–722 (2019).
The innate immune system is the first line of defense against bacterial and viral infection. Pathogen-derived nucleic acids such as DNA and RNA induce potent immune responses in the infected cells.
The detection of viral nucleic acids is an important strategy by which the host senses infection and initiates protective immune responses. The cGAS-STING pathway plays a central role in the innate immune response towards viral infection. Cyclic GMP-AMP synthase (cGAS) is activated by viral DNA and catalyzes the synthesis of a cyclic dinucleotide called cGAMP, which stimulates the induction of interferons through the STING-TBK1-IRF3 signaling axis.
We use biochemical, biophysical, structural, and cellular approaches to elucidate the molecular mechanisms of viral DNA sensing through the cGAS-STING pathway. These studies will advance our understanding of the molecular bases of innate immunity against viral infection and provide a structural framework for innovative approaches to treat viral disease, autoimmune disorder, and cancer.
Publications
- View publications on Google Scholar