Department of
Biochemistry and Biophysics

9+ Research Areas
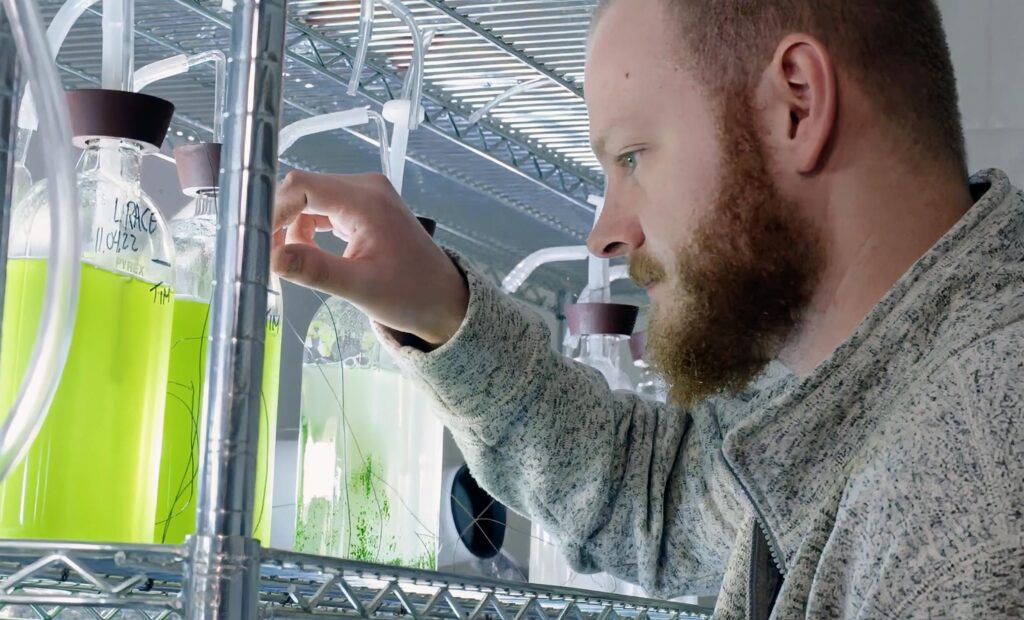
Our department is at the forefront of cutting-edge research, delving into the intricate molecular mechanisms that underpin life itself. Our scientists explore diverse scientific problems across the fields of biochemistry, biophysics, genetics, molecular biology, and genomics.

38+ Research-Active Faculty
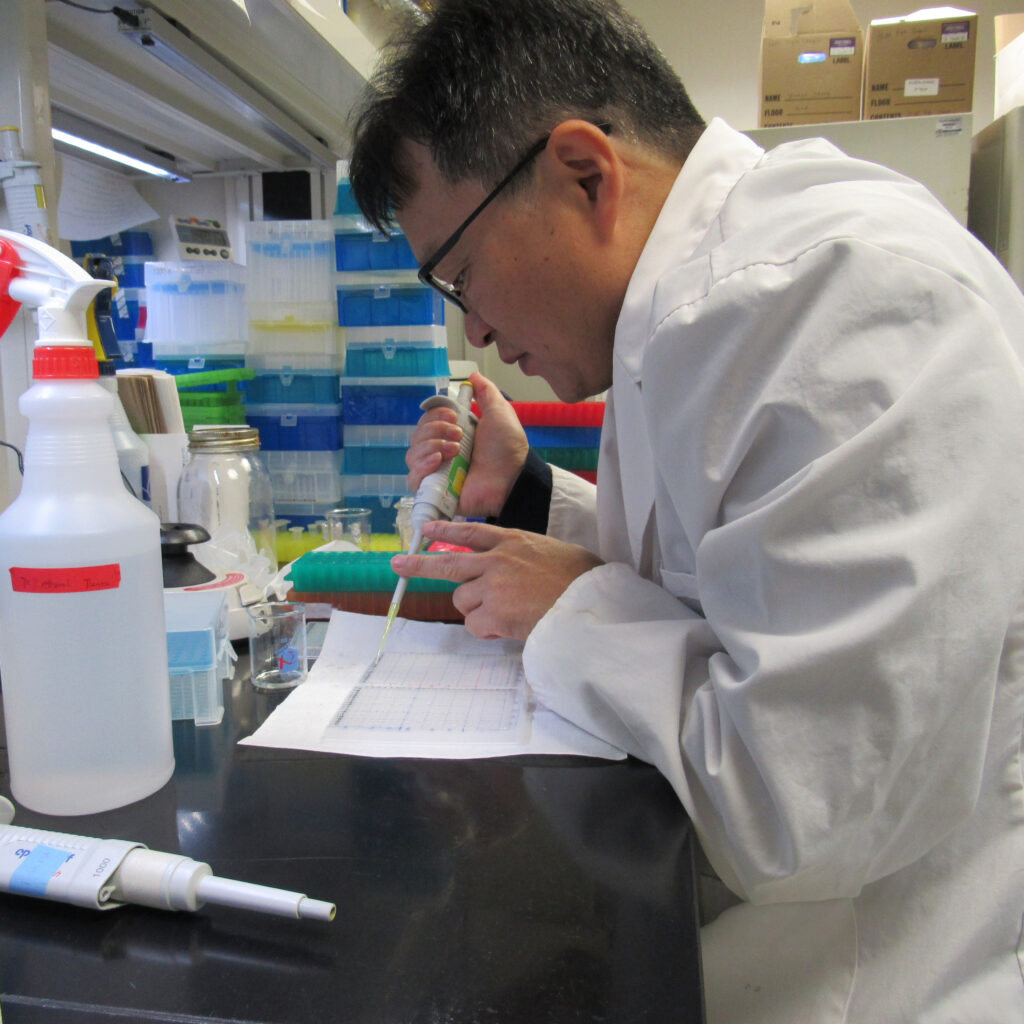
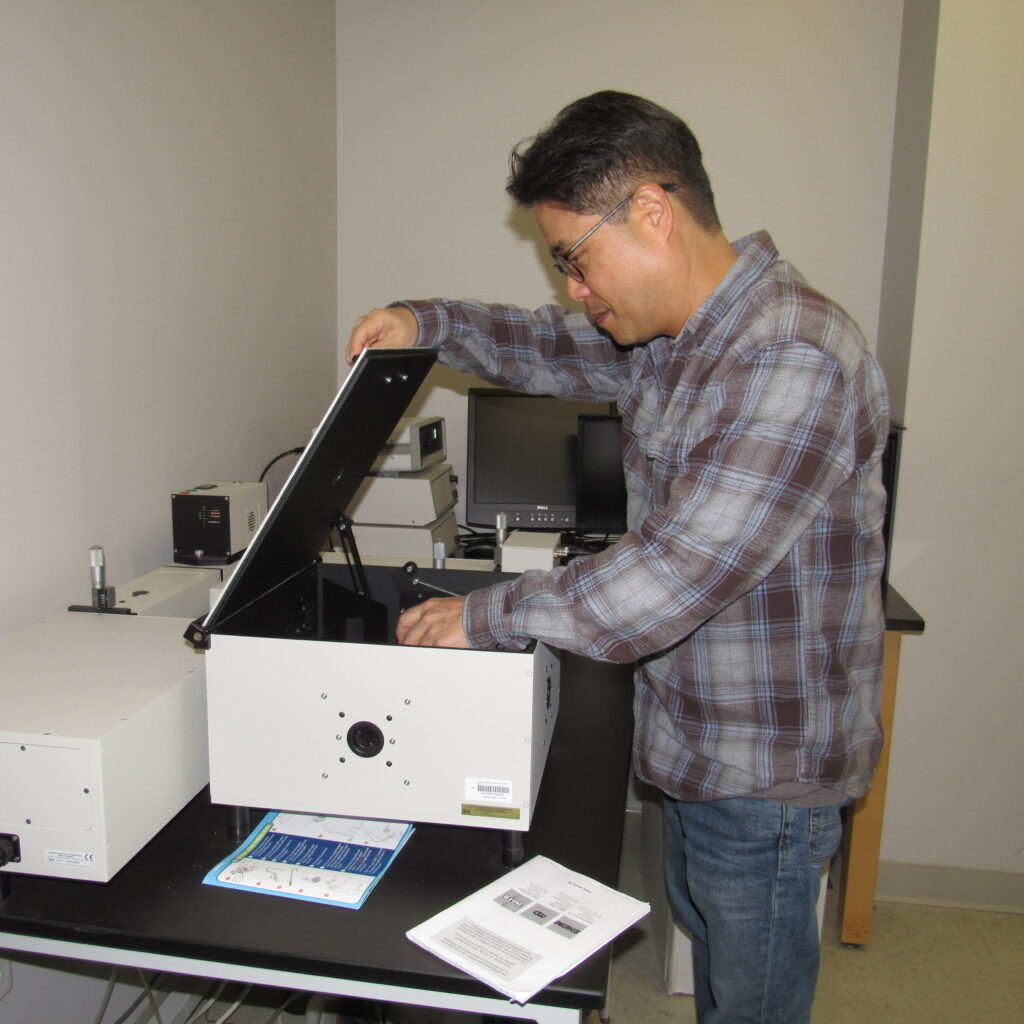
Our research-active faculty engage in investigating practical applications to address real-world issues, simultaneously striving to push the boundaries of interdisciplinary paradigms and mentor the next generation of biochemical and biophysical scientists.

600+ Enrolled Students
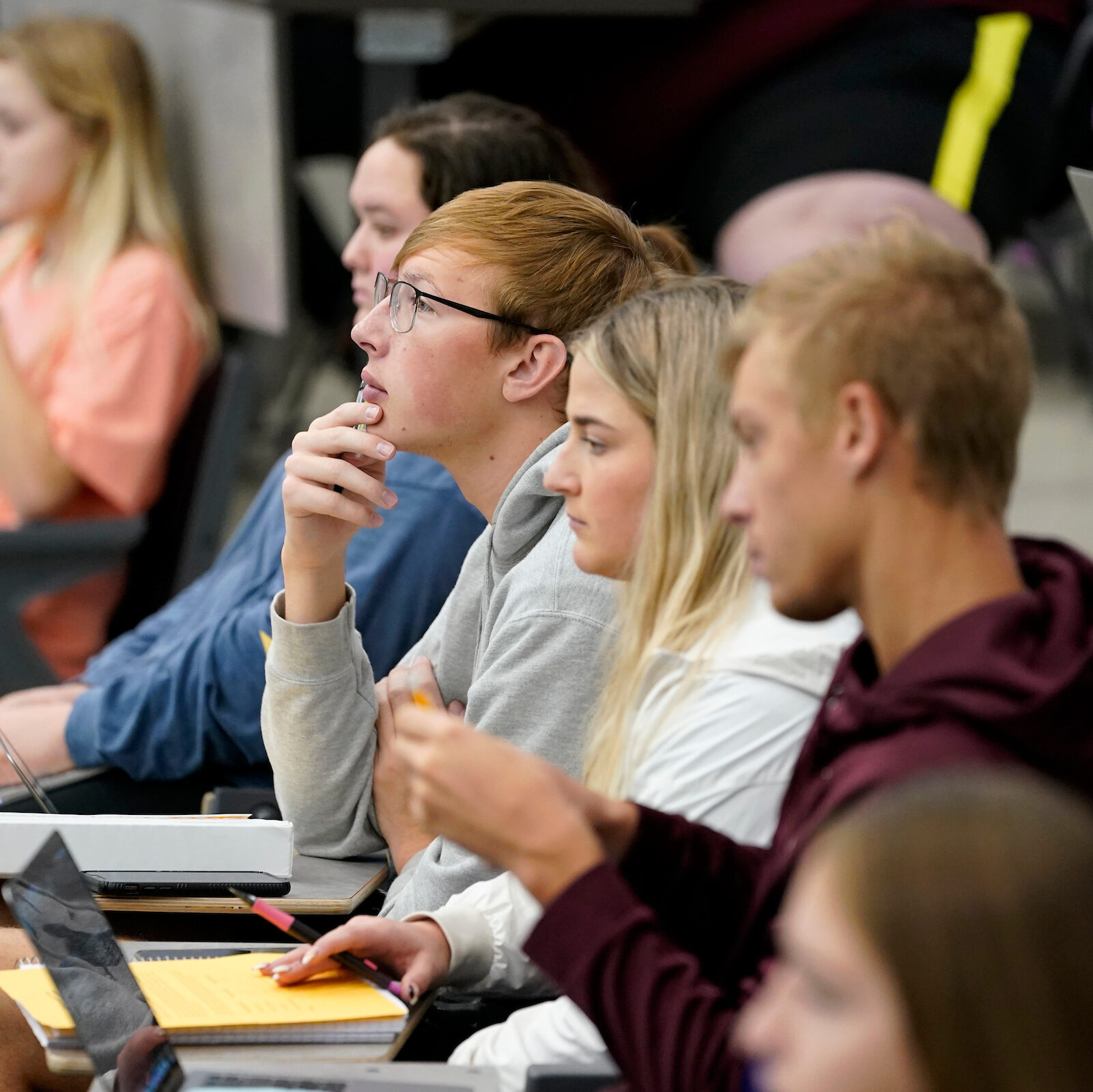
Our undergraduate and graduate students learn both in the classroom and through hands-on research experiences, developing unique skillsets required for success in academia, professional school, or a career in scientific industry.

Recent Publications
Department Updates
-

“Ms. Harman Kaur, a graduate student in the Gohil lab, won the outstanding presentation award at the 2024 Texas Genetics Society Meeting.”
A little bit about Ms. Kaur, her major is Biochemistry. Her class year is 2026. Also, she is a departmental representative of WISE (Women in…
-
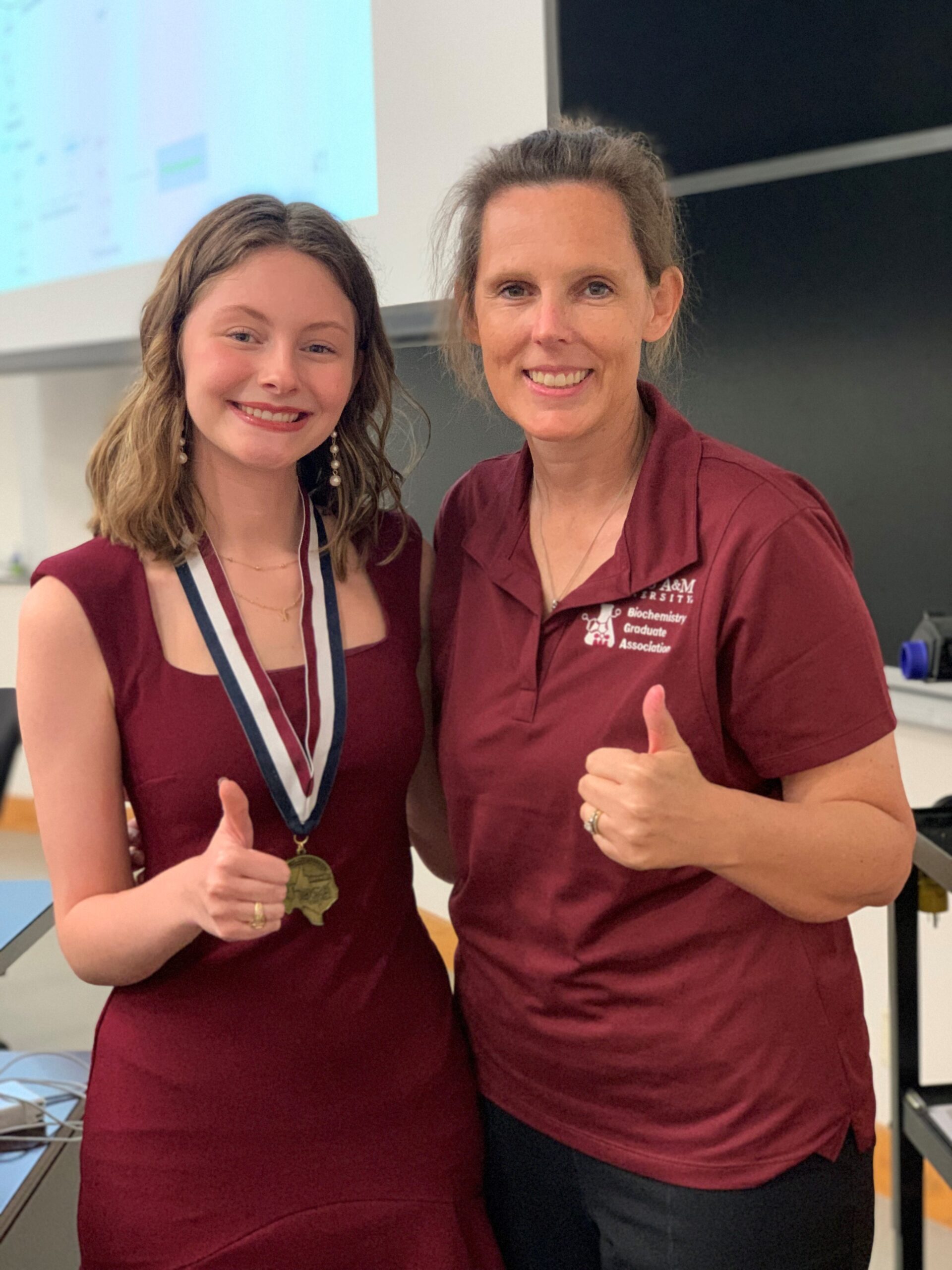

Congratulations Katelyn Willis
Junior neuroscience major Katelyn Willis recently placed first in an exam-based biochemistry competition against other undergraduate students at the Texas HOSA Spring Leadership Conference. The annual…
-
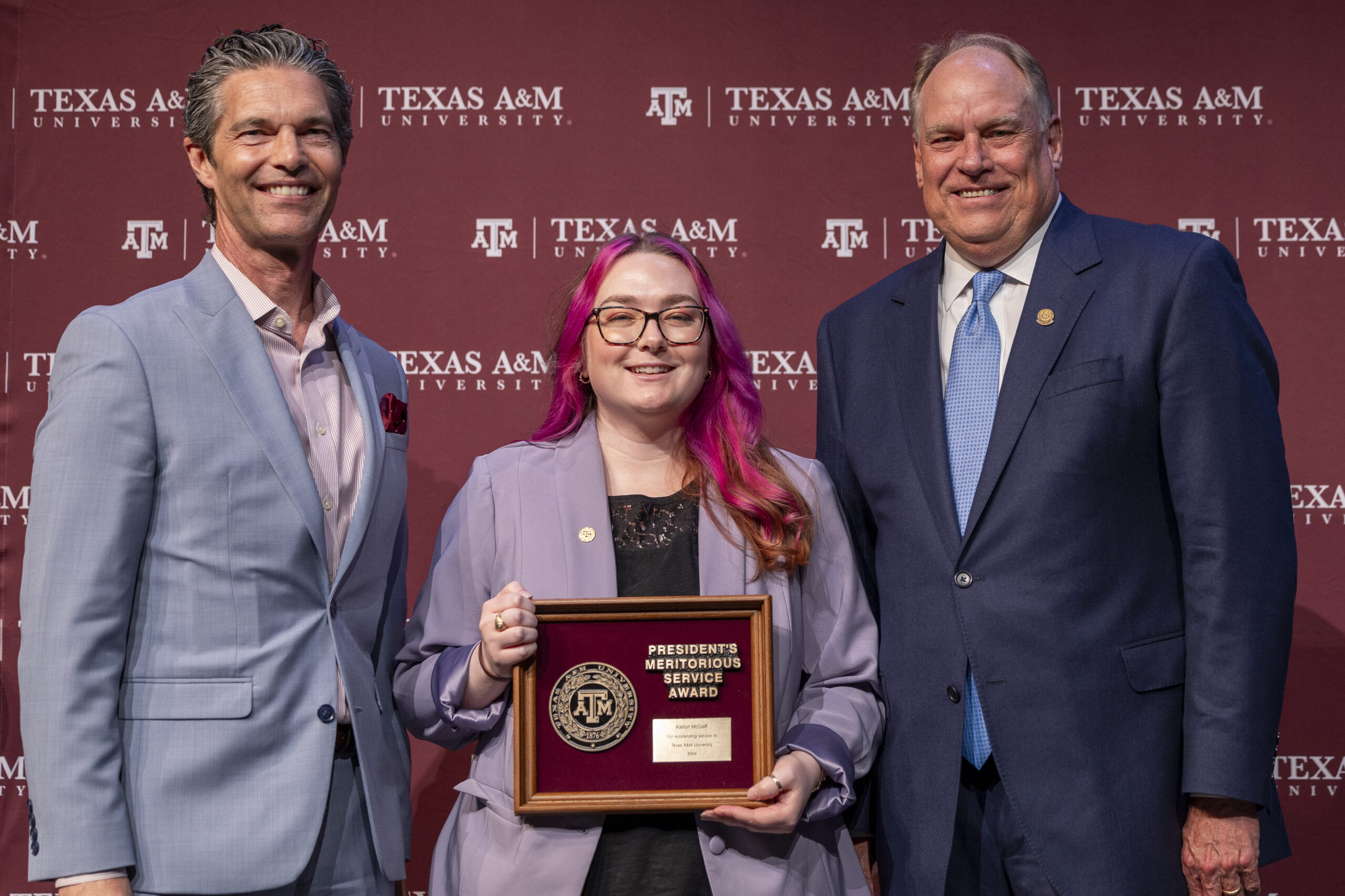

Congratulations to Kaitlyn McGaff on receiving the President’s Meritorious Service Award.
This Award is given to individuals who demonstrate their commitment to the Aggie Core Values of Respect, Excellence, Leadership, Loyalty, Integrity and Selfless Service. Way…
Biochemistry and Biophysics News

Researchers resolve old mystery of how phages disarm pathogenic bacteria
Bacterial infections pose significant challenges to agriculture and medicine, especially as cases of antibiotic-resistant bacteria continue to rise. In response, scientists at Texas A&M AgriLife Research are elucidating the ways that bacteria-infecting viruses disarm these pathogens and ushering in the possibility of novel treatment methods.

Impacting Texans’ lives as a neurosurgeon, legislator and medical volunteer
When State Rep. James “Greg” Bonnen ’88, M.D., arrived to deliver a guest lecture at the Texas A&M College of Agriculture and Life Sciences Department of Biochemistry and Biophysics on Feb. 22, it was a customary occasion for the accomplished neurosurgeon, legislator and loyal former student.
Have Questions?
For undergraduate admissions questions:
For graduate admissions questions:
For general questions: